Home  Introduction Introduction  Overview Overview |
||||
See also: Application Example: Bank Card, Application Example: Elemental Analysis of Concrete, The Data Cube, Image Stack, Application Example: Enrichment in a Dried Droplet, Application Example: Multisensor Analysis of Particulate Matter, Metadata, Workflow
 |
||||
Overview |
||||
|
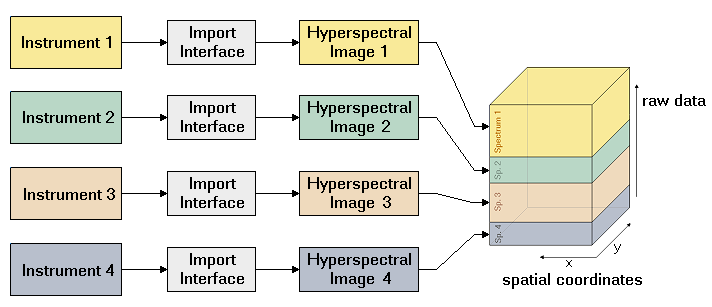
Epina ImageLab is a Windows-based multisensor imaging system for processing and analyzing hyperspectral images. It is a modular system consisting of a basic engine, a graphical user interface, and optional user-supplied modules. It supports all major spectroscopic imaging techniques, such as UV/VIS/NIR, infrared, Raman, THz, optical emission and absorption, EDX (energy dispersive xray), and mass spectrometry. Epina ImageLab also supports the combination of up to four spectroscopic techniques, allowing to intertwine the data measured by different instruments to a single data set. Thus complementary information from different spectroscopic techniques can be exploited to gain additional insight into complex samples.
A plethora of built-in multivariate statistical methods enables the user to analyze and classify acquired hyperspectral images. Currently available multivariate techniques are:
All images - either deriving from raw data or statistical analyses - can be combined with conventional photos, thus creating an image stack consisting of up to eight layers. Using different algorithms and levels of transparency analytical information can be highlighted directly in the corresponding photo, making it much easier to recognize areas of interest. As an aid to the researcher, Epina ImageLab provides several tools to preprocess and improve raw image data. If necessary images can be cut, resampled, mirrored and masked. A bad pixel finder helps to identify pixels which are invalid due to problems with the detector. Furthermore there are several tools which can be used to process the spectral information. The most important ones help you with baseline subtraction, smoothing, and the calculation of derivative spectra. One particular strength of Epina ImageLab is based on so-called spectral descriptors, which help to introduce chemical and physical knowledge into the statistical analysis of the images. By defining spectral descriptors the multidimensional space describing the samples is augmented by additional variables. It has been shown that the classification of images becomes considerably more reliable by using spectral descriptors.
In addition to many other imaging software packages, Epina ImageLab also supports the time domain as a fourth dimension. Thus the user may visualize data over time, creating a video highlighting changes in the sample (provided that the sample can be measured without destruction). Epina ImageLab provides a versatile and open programming interface allowing researchers to hook up their own data processing modules. A built-in scripting engine allows to automate daily tasks and can be used to write customized image analysis routines. In addition, the concept of user-defined modules can be used to develop data import modules for any spectroscopic device.
|
||||